Plot model fits (observed and fitted) and the residuals for a row or column of an mmkin object
Source:R/plot.mmkin.R
plot.mmkin.RdWhen x is a row selected from an mmkin object ([.mmkin), the
same model fitted for at least one dataset is shown. When it is a column,
the fit of at least one model to the same dataset is shown.
Arguments
- x
An object of class
mmkin, with either one row or one column.- main
The main title placed on the outer margin of the plot.
- legends
An index for the fits for which legends should be shown.
- resplot
Should the residuals plotted against time, using
mkinresplot, or as squared residuals against predicted values, with the error model, usingmkinerrplot.- ylab
Label for the y axis.
- standardized
Should the residuals be standardized? This option is passed to
mkinresplot, it only takes effect ifresplot = "time".- show_errmin
Should the chi2 error level be shown on top of the plots to the left?
- errmin_var
The variable for which the FOCUS chi2 error value should be shown.
- errmin_digits
The number of significant digits for rounding the FOCUS chi2 error percentage.
- cex
Passed to the plot functions and
mtext.- rel.height.middle
The relative height of the middle plot, if more than two rows of plots are shown.
- ymax
Maximum y axis value for
plot.mkinfit.- ...
Further arguments passed to
plot.mkinfitandmkinresplot.
Details
If the current plot device is a tikz device, then
latex is being used for the formatting of the chi2 error level.
Examples
# \dontrun{
# Only use one core not to offend CRAN checks
fits <- mmkin(c("FOMC", "HS"),
list("FOCUS B" = FOCUS_2006_B, "FOCUS C" = FOCUS_2006_C), # named list for titles
cores = 1, quiet = TRUE, error_model = "tc")
#> Warning: Optimisation did not converge:
#> iteration limit reached without convergence (10)
plot(fits[, "FOCUS C"])
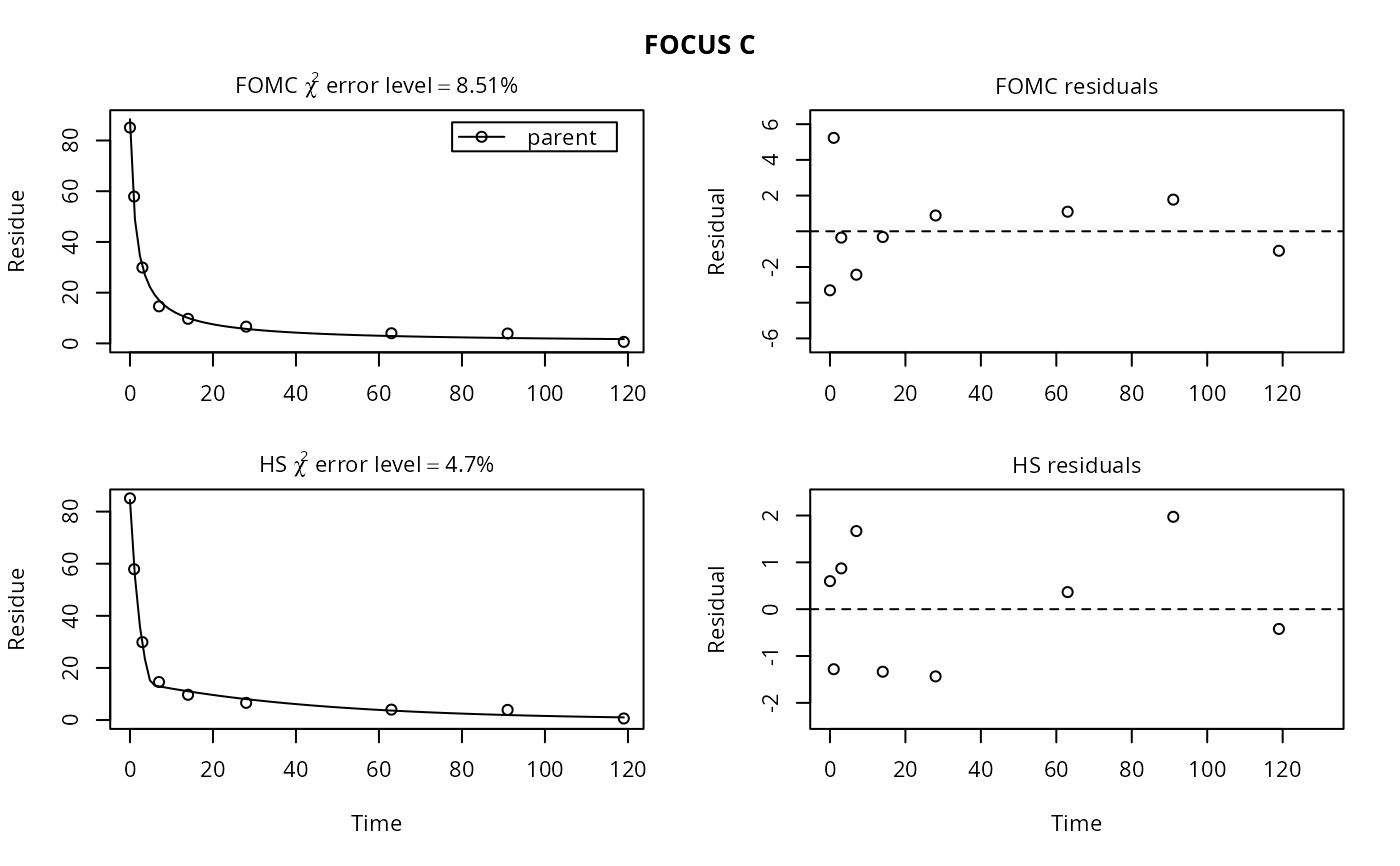 plot(fits["FOMC", ])
plot(fits["FOMC", ])
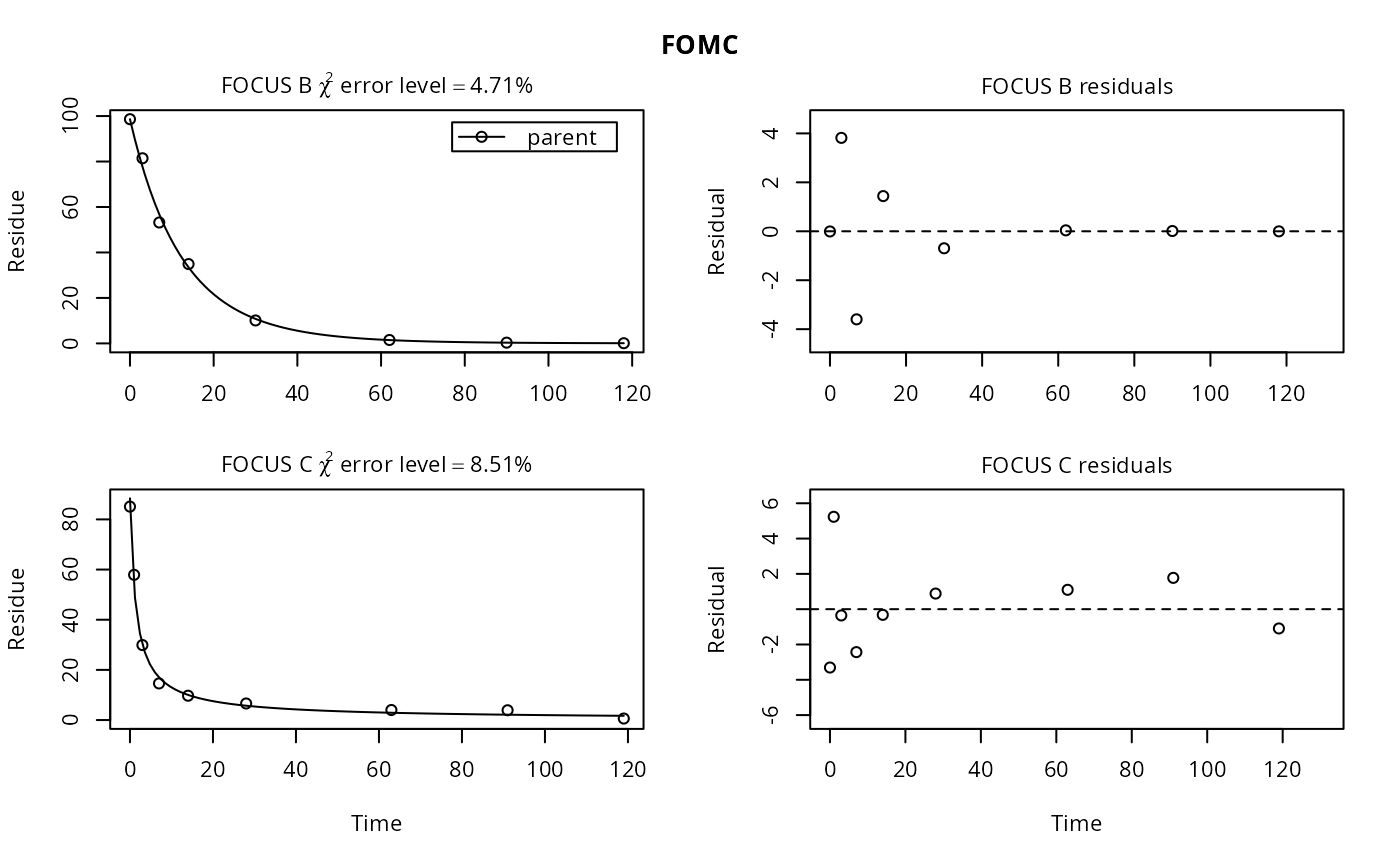 plot(fits["FOMC", ], show_errmin = FALSE)
plot(fits["FOMC", ], show_errmin = FALSE)
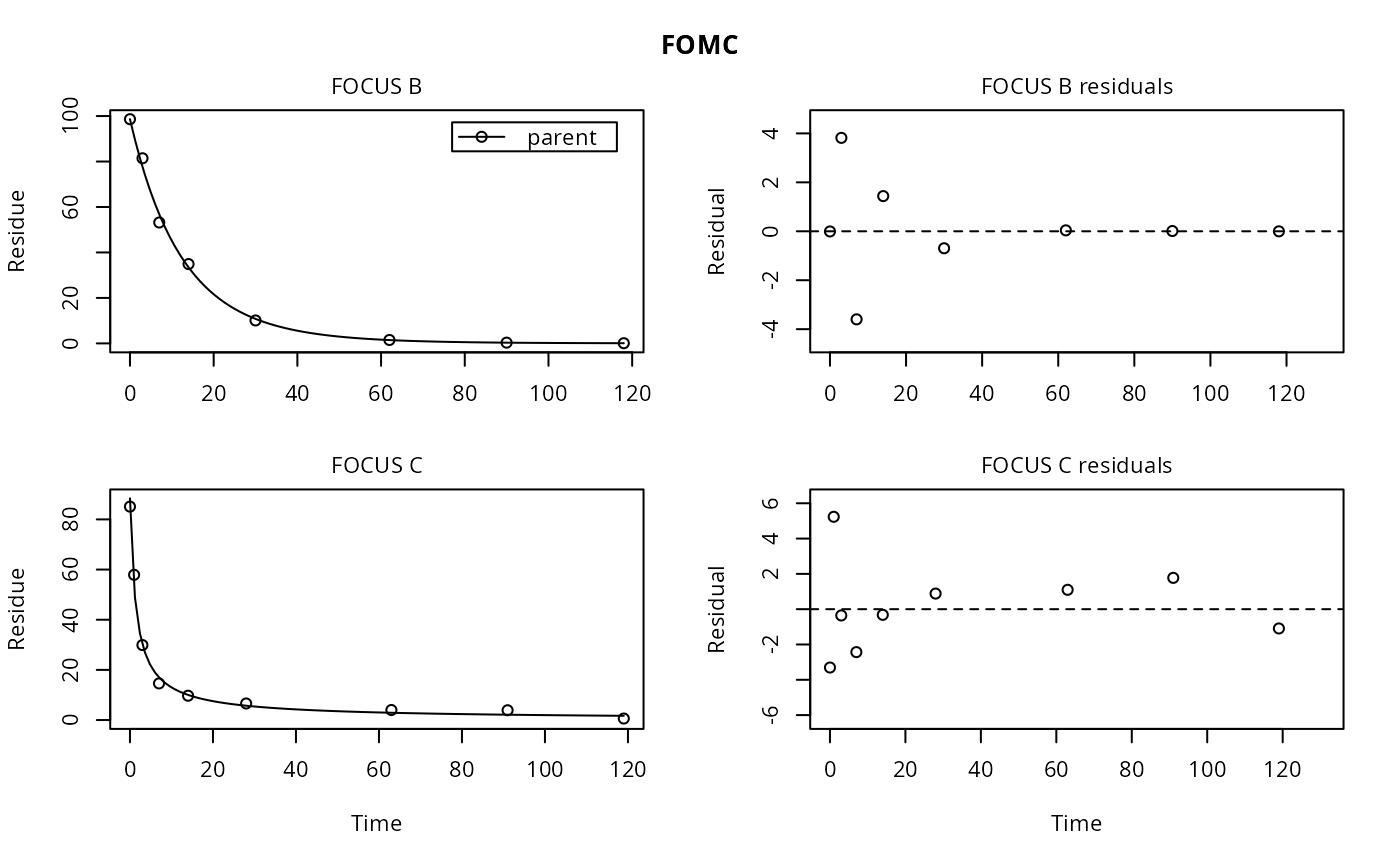 # We can also plot a single fit, if we like the way plot.mmkin works, but then the plot
# height should be smaller than the plot width (this is not possible for the html pages
# generated by pkgdown, as far as I know).
plot(fits["FOMC", "FOCUS C"]) # same as plot(fits[1, 2])
# We can also plot a single fit, if we like the way plot.mmkin works, but then the plot
# height should be smaller than the plot width (this is not possible for the html pages
# generated by pkgdown, as far as I know).
plot(fits["FOMC", "FOCUS C"]) # same as plot(fits[1, 2])
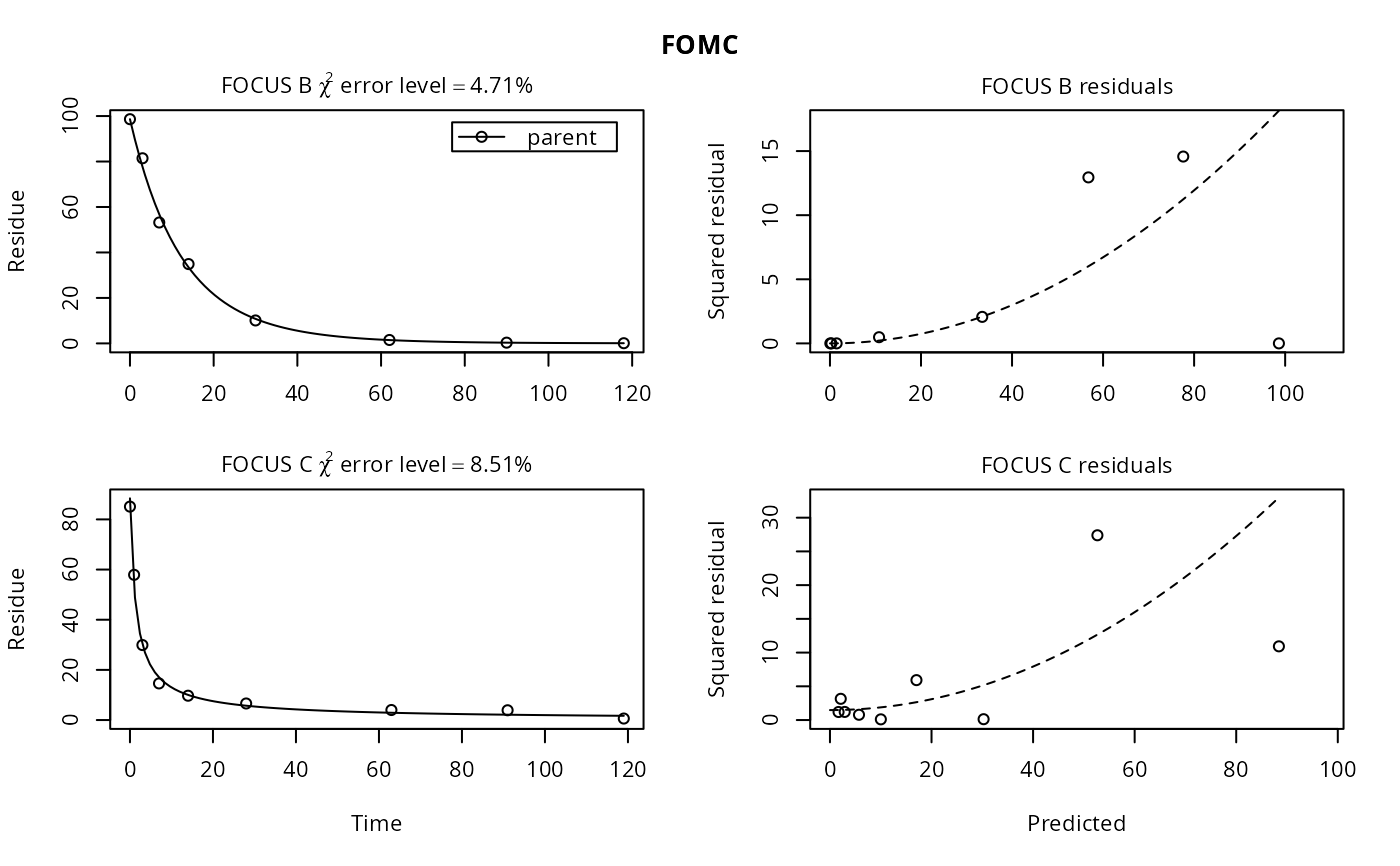
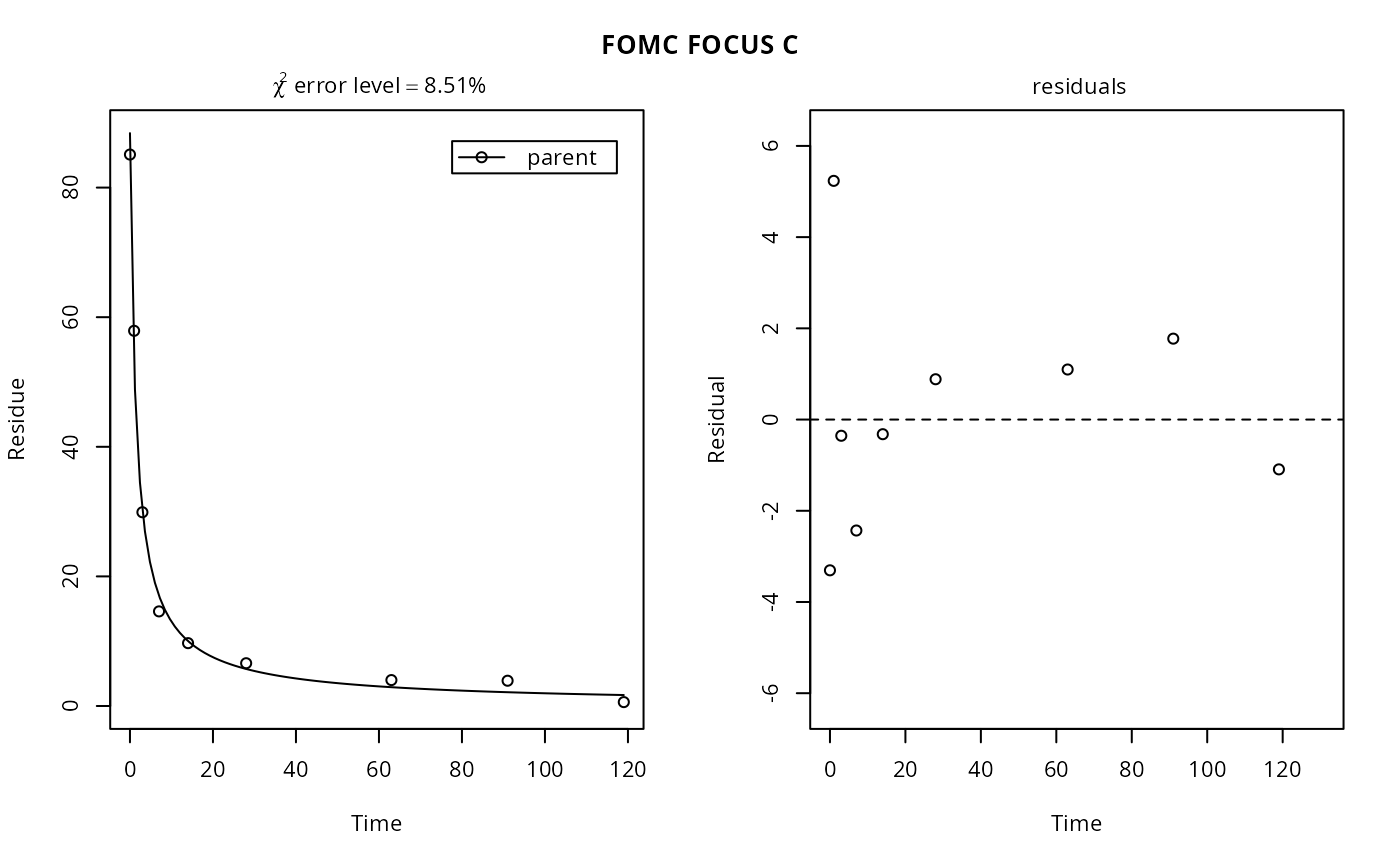 # Show the error models
plot(fits["FOMC", ], resplot = "errmod")
# Show the error models
plot(fits["FOMC", ], resplot = "errmod")
 # }
# }