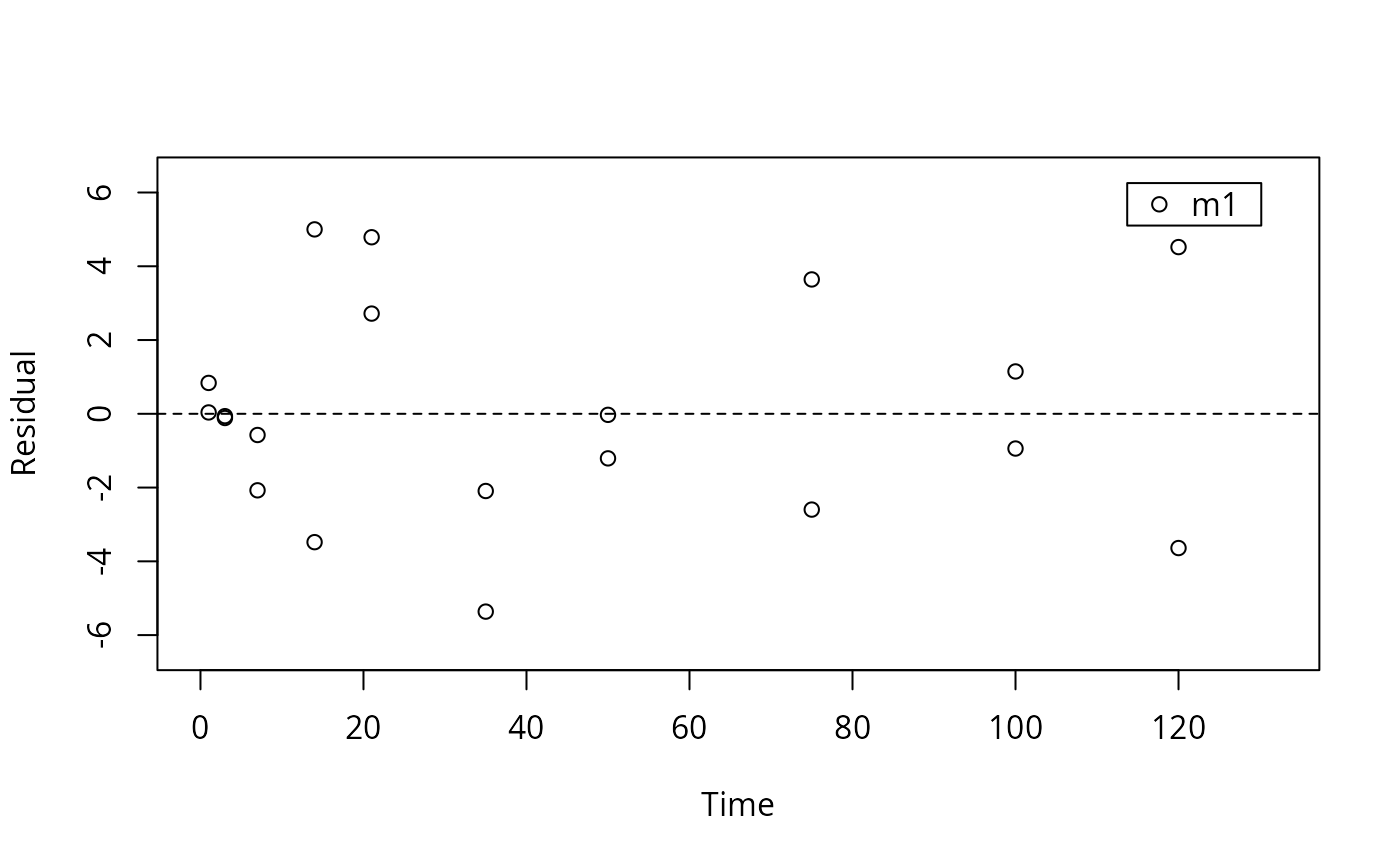
This function plots the residuals for the specified subset of the observed
variables from an mkinfit object. A combined plot of the fitted model and
the residuals can be obtained using plot.mkinfit using the
argument show_residuals = TRUE.
Usage
mkinresplot(
object,
obs_vars = names(object$mkinmod$map),
xlim = c(0, 1.1 * max(object$data$time)),
standardized = FALSE,
xlab = "Time",
ylab = ifelse(standardized, "Standardized residual", "Residual"),
maxabs = "auto",
legend = TRUE,
lpos = "topright",
col_obs = "auto",
pch_obs = "auto",
frame = TRUE,
...
)Arguments
- object
A fit represented in an
mkinfitobject.- obs_vars
A character vector of names of the observed variables for which residuals should be plotted. Defaults to all observed variables in the model
- xlim
plot range in x direction.
- standardized
Should the residuals be standardized by dividing by the standard deviation given by the error model of the fit?
- xlab
Label for the x axis.
- ylab
Label for the y axis.
- maxabs
Maximum absolute value of the residuals. This is used for the scaling of the y axis and defaults to "auto".
- legend
Should a legend be plotted?
- lpos
Where should the legend be placed? Default is "topright". Will be passed on to
legend.- col_obs
Colors for the observed variables.
- pch_obs
Symbols to be used for the observed variables.
- frame
Should a frame be drawn around the plots?
- ...
further arguments passed to
plot.
Value
Nothing is returned by this function, as it is called for its side effect, namely to produce a plot.